Tutorial
This short tutorial will show you how to use GenExp to explore genomes and will highlight some of its features.
Starting with GenExp
GenExp is a web-based application and so it's not necessary to download or install anything. To use it, simply connect to https://gralggen.lsi.upc.edu/recerca/genexp using your web-browser and you will see something like this:
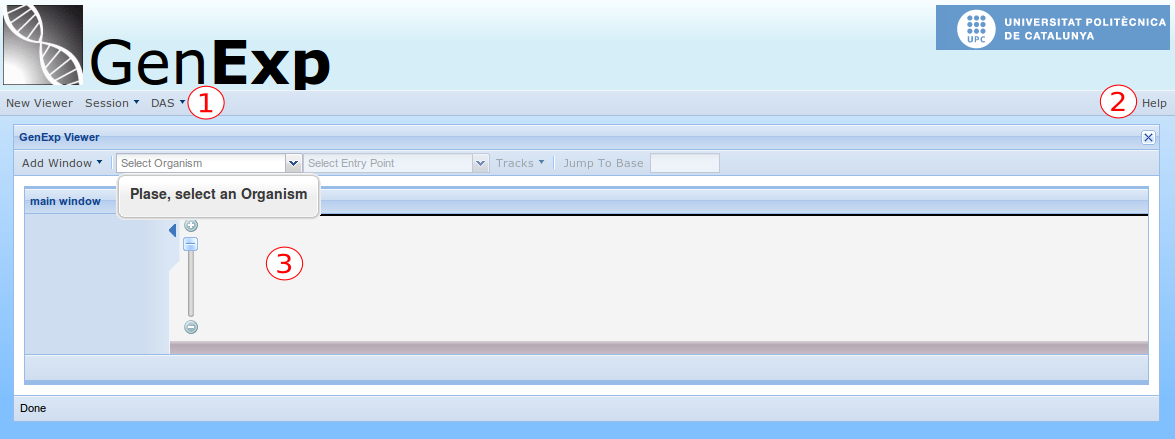
This is the main view of GenExp. You can see the Main Toolbar (1) where the buttons and menus affecting the whole system are. From that toolbar you can add and remove data sources and save and load browsing sessions. At the far right side of the main toolbar, the Help button (2) lives. This button is always there in case you need some more information on anything regarding GenExp.
The blank frame just below the main toolbar is a Viewer (3). This is the key component of GenExp and is where the genome data will be drawn. Each viewer has its own toolbar with all the actions affecting only that specific viewer, such as changing the organism or chromosome, and selecting the data sources to be drawn
Selecting organism and chromosome
The viewer's toolbar is used to select what region do we want to show. We will first select the organism. To do that, click on the organism combobox (1) and select "Homo sapiens - GRCh37". The Entry Point selector will become active. The Entry Point selector is used to select among the sequences on the organism (in this case, basically the chromosomes). Select chromosome 2 and the Tracks menu will become active.
At this point you should see something like this
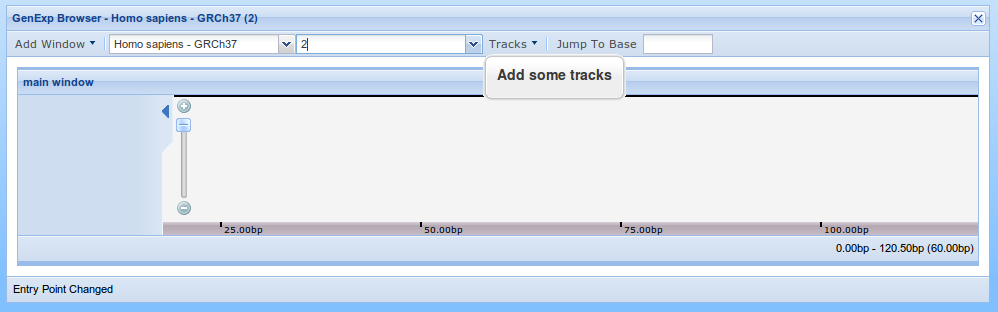
Adding the reference sequence
For every organism listed there's a special data track to add the reference genomic sequence. All data sources are listed under the Tracks menu. Click on it and select "ensembl GRCh37 Reference" and then "Reference Sequence".
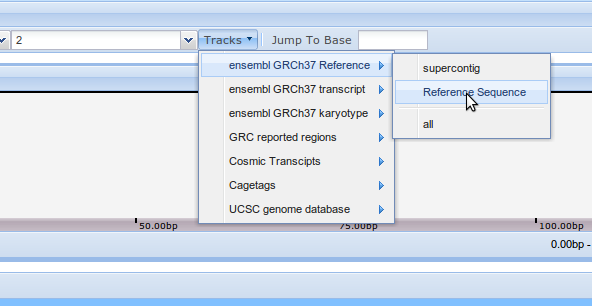
A list of n's will appear on the viewer bottom. This is because the first part of the sequence has not been exactly determined, yet, and it's represented by n's.
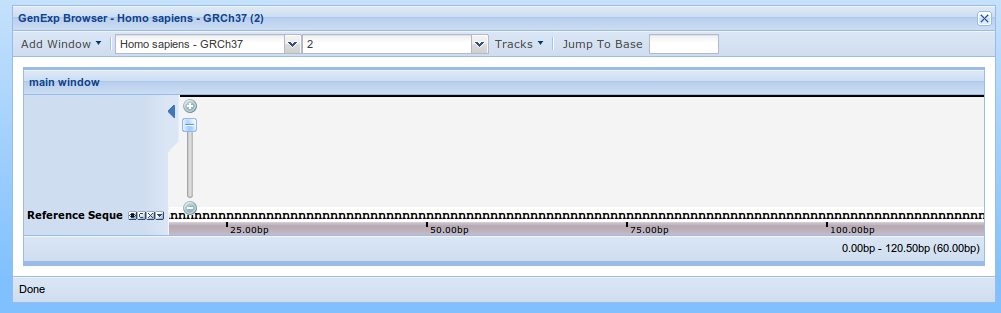
Adding data
Although data from any DAS source can be added, a preselected list of data sources has been added. Click on the Tracks button to open the list preselected data sources. Each of those sources has one or more types of data available. Click on "ensembl GRCh37 transcript" and select "exon:coding". A new track will appear over the Reference sequence with the coding exons from ensembl. Do the same to add "cosmic:exon" from "Cosmic Transcripts". A track with the exons on COSMIC database will be created. Finally add "all" from "GRC reported regions", the in the current assembly reported to the GRC.
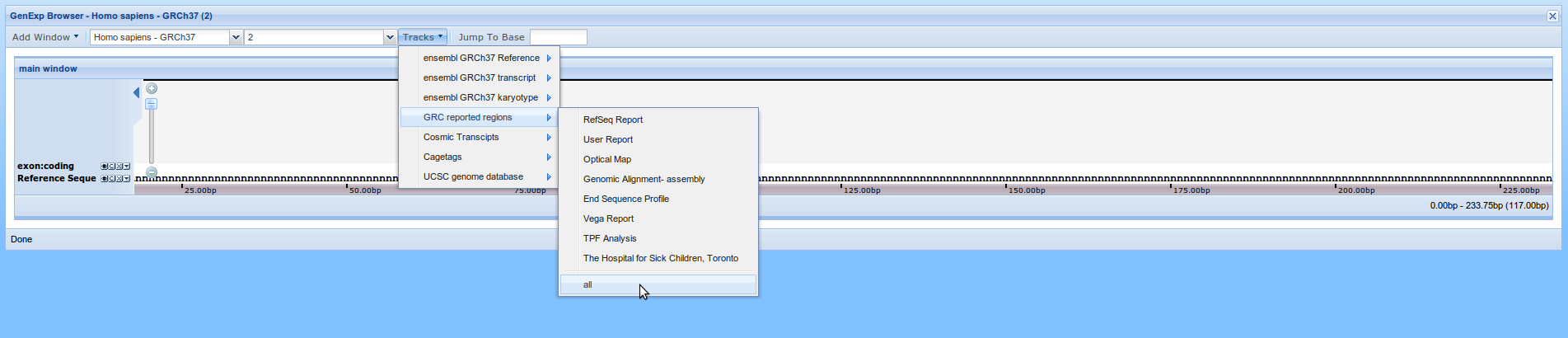
As you can see here, nothing appeared on the viewer. This is because there's nothing on this region. As you can see at the bottom right corner of the viewer, the represented region is from base 0 to 233 and centered at 117. This means we are seeing the first 233 bases of the chromosome and there's nothing here.
Moving around
To get a more interesting view, we'll change the region being drawn. To change the zoom you can use the zoom slider (1) or the mouse wheel as in any map application. Click on plus or minus, drag the slider or use the mouse wheel to change the zoom level. At some point you will see some features appear.

Note:You will also see that while most of the zoom changes are done immediately, in some cases gray dots will cover the tracks while data is loaded. This happens because the system needs the data in a different resolution.
To move around the chromosome you can drag the image with your mouse. Click anywhere on the viewer canvas (the gray/white space where the features are drawn) and move the mouse without releasing the button. The image will move and new parts of the chromosome will be revealed. To move to an exact place you can use the "Jump to base" functionality.
On the top of the viewer, next to the "Jump to Base" button, write 110000000 and click on the "Jump to Base" button. The viewer will now be centered at 110Mb. Use the zoom to get a view like that.
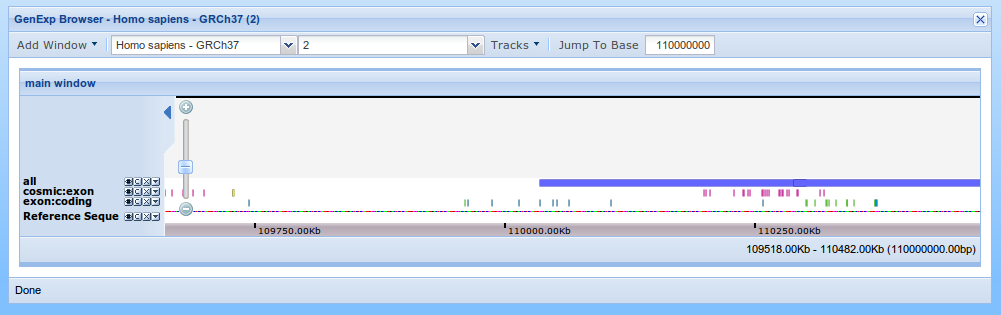
If you want to see an entire feature on the viewer, you can move the mouse over the feature and on the pop up message. click on the "Show Feature" button. If the feature is partially occluded by the track controls, click on the "hide controls" button to hide them.

Loading Data
At some moments GenExp will need to load new data from the data source. In that case, a gray area will cover the tracks and a small label will report some information about the data loading procedure. On that label you will see a message reading "Loading...", a couple of numbers X/Y stating that Y packages of data need to be loaded and Y have been loaded so far, and an estimate of the maximum time it will take to load the data on that track.
Note:Please, bear in mind that the time is an estimate and that a data package can take longer than expected to load, specially when loading very large portions of a chromosome. In some cases, loading data from slow data source may take up to a few minutes.
If at some point a package take much more than expected to load the label will change its color to warn of a possible network problem. However, GenExp will keep trying to fully load the requested data.
Multiple Windows
It is possible to add multiple windows of the same data via the addition of dependent windows. There are two types of dependent windows: "Overview" windows and "Next/Previous" windows.
"Overview" windows are show the same region as the main window but at an independent zoom level. To create one of those windows, click on "Add Window" and "Overview".
A new window will appear inside the viewer with a lower zoom level. You can change the zoom levels of the two windows independently but they will always synchronized (i.e. both windows will be centered on the same base of the same sequence). Try moving one of them and the other will move accordingly.

With "Add Window" "Previous" and "Next", a new window will be added to the viewer showing the region next to the main window. For example, if the main window shows the region from 0 to 100 bases, the "Next" will show from "100 to 200". Add a "Next" window and you will end up with something like this.
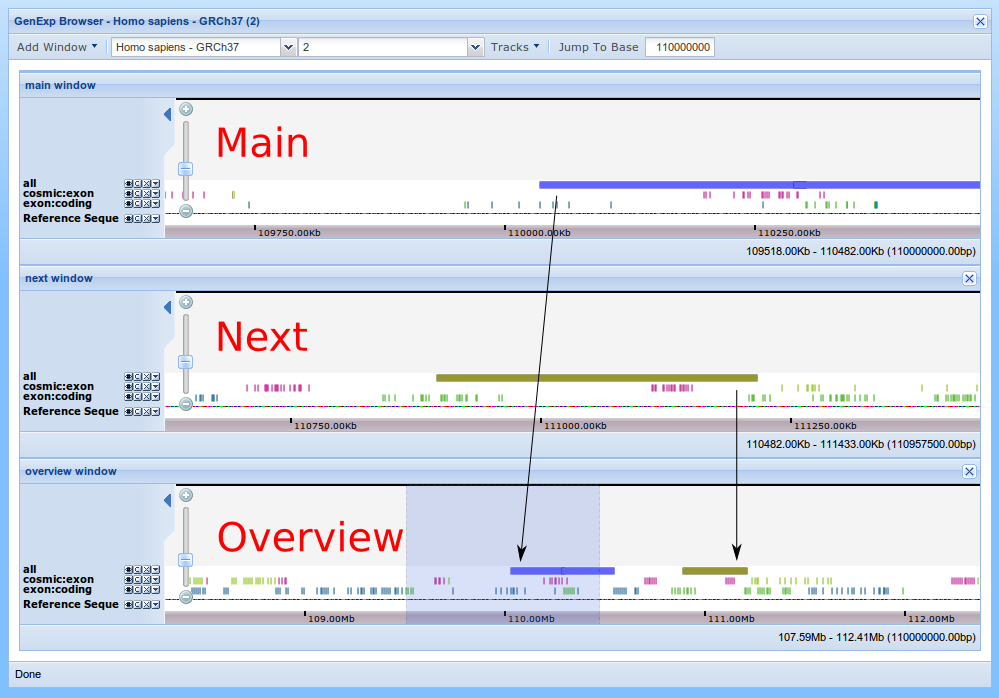
Any of those windows can be closed clicking on the small cross at its top-right corner.
Multiple Views
It is also possible to add independent viewers to the same page. Those independent viewers can represent completely independent data to compare it or even the same data is desired.
To add a new viewer click on the "New Viewer" on the main toolbar. For example, after closing the dependent windows and adding a new viewer showing human chromosome 17, the application looks like this.
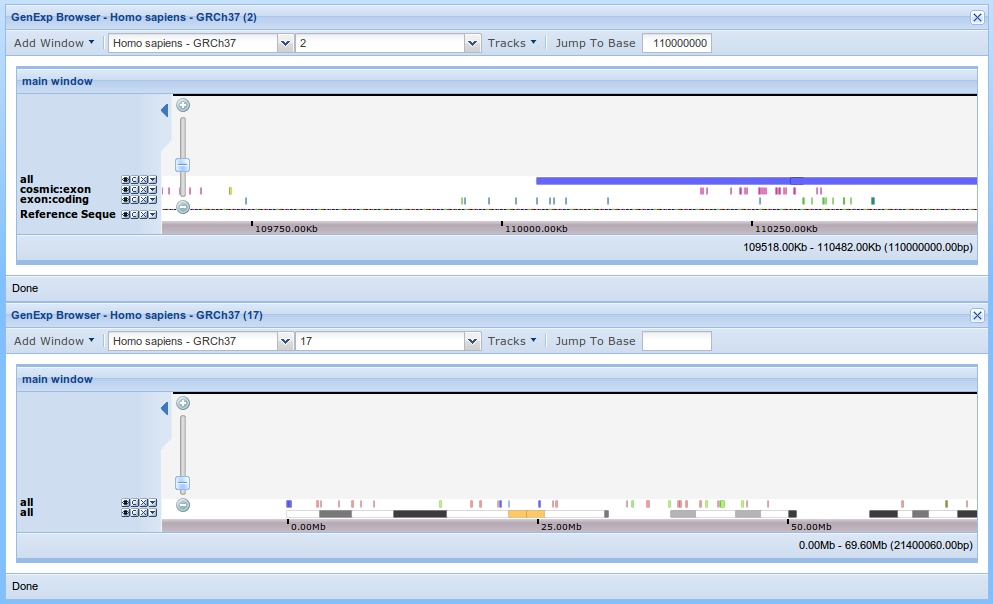
Additional functionality
Track Controls
Track controls are the small buttons next to each track. You can hide and remove tracks, change its appearance and change its order. When moving and zooming out a lot, hiding very dense or slow tracks can improve performance. Once you unhide them, data retrieval will start automatically.
Sessions
It is possible to store a browsing session and reuse it later. To do it use the "Session" menu on the main toolbar.
Custom Sources
Although only some data sources have been preselected any DAS source can be added to GenExp. Click on "DAS"->"Add DAS Source" on the main menu, enter the URL of a valid DAS source (for example your own DAS source on easyDAS) and give it a name. It will be automatically added to the system and available on the "Tracks" menu on the viewers. Custom DAS sources are not stored on the server and only used on the client side.